Creating meshless domains¶
In this tutorial we will see how to:
[1]:
# import necessary libraries
import matplotlib.pyplot as plt
import torch
Some general geometries in 1D, 2D and 3D are already implemented.
[2]:
# import classical domains
from scimba_torch.domain.meshless_domain import (
Cube3D,
Cylinder3D,
Disk2D,
Disk3D,
Polygon2D,
Segment1D,
Square2D,
Torus3D,
TorusFrom2DVolume,
)
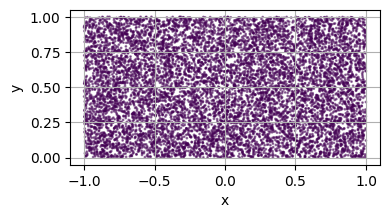
Creating a simple Domain¶
To create a simple domain, simply instantiate one of the meshless_domain class with the appropriate parameters.
is_main_domain is used to specify if the domain can have subdomains/holes.[3]:
square = Square2D(
bounds=[(-1, 1), (0, 1)],
is_main_domain=True
)
# to reset the square domain
def simple_square():
return Square2D(
bounds=[(-1, 1), (0, 1)],
is_main_domain=True
)
To visualise a domain, we are going to sample points on it
[4]:
# import necessary libraries
from scimba_torch.integration.monte_carlo import DomainSampler
# a utility function to plot a 2D domain
def plot_2d(domain, n=10_000, ax=None):
"""Plot a 2D domain using a Domain sampler."""
# create the sampler
sampler = DomainSampler(domain)
# sample points
res = sampler.sample(n)
pts = res.x # coordinates of the sampled points
labels = res.labels # labels of the sampled points
# convert to numpy for plotting
pts = pts.detach().cpu().numpy()
labels = labels.detach().cpu().numpy()
# generate the ax if not provided
if ax is None:
fig = plt.figure(figsize=(4, 4))
ax = fig.add_subplot(111, xlabel='x', ylabel='y', aspect='equal')
else:
fig = None
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_aspect('equal')
# plot the points
ax.scatter(pts[:, 0], pts[:, 1], s=1, c=labels, alpha=0.5)
ax.grid(True)
if fig is not None:
fig.tight_layout()
plt.show()
square = simple_square()
plot_2d(square)
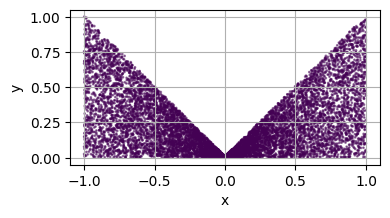
Apply a Mapping to a Domain¶
For more informations about mappings, see Complement on the mapping.
[5]:
from scimba_torch.utils.mapping import Mapping
# define a mapping function
def map(x):
"""A simple mapping function."""
# x: (N, 2) tensor
# out: (N, 2) transformed tensor
return torch.stack([x[:, 0], x[:, 1]*(torch.abs(x[:, 0]) + 1e-2)], dim=1)
# define the mapping
mapping = Mapping(
from_dim=2, # original dimension
to_dim=2, # mapped dimension
map=map, # mapping function
inv=None, # optional inverse mapping function
)
# set the mapping for the square domain
square = simple_square()
square.set_mapping(
map=mapping,
bounds_postmap=[(-1, 1), (0, 1)], # bounding box of the MAPPED domain
)
plot_2d(square)
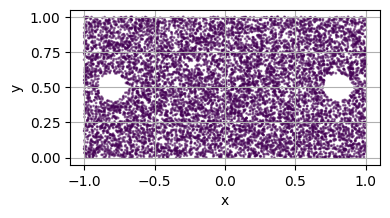
Create a Domain with holes¶
is_main_domain set to false.For example, let’s consider a disk hole inside the domain
[6]:
square = simple_square()
# create some holes
hole1 = Disk2D(center=(.8, 0.5), radius=0.1, is_main_domain=False)
hole2 = Disk2D(center=(-.8, 0.5), radius=0.1, is_main_domain=False)
# add holes to the square domain
square.add_hole(hole1)
square.add_hole(hole2)
plot_2d(square)
[7]:
# helper for resetting the domains
def simple_disks():
disk_L = Disk2D(center=(-.8, 0.5), radius=0.1, is_main_domain=False)
disk_R = Disk2D(center=(.8, 0.5), radius=0.1, is_main_domain=False)
return disk_L, disk_R
Combining Holes and Mapping¶
We can also combine holes and mappings.
adding a hole (if the mapping is already set), with the flag
copy_mappingset to Falsesetting the mapping for all already added holes, with the flag
to_holesset to False
Yet, we recommend to proceed in the following order:
Create the main domain
Set the mapping if any
Add holes to the domain, with
copy_mappingset to False if we do not want to apply the mapping on the hole
[8]:
fig, axs = plt.subplots(1, 3, figsize=(12, 4), sharex=True, sharey=True)
# by default, the mapping is applied to all the holes
square = simple_square()
disk_L, disk_R = simple_disks()
square.set_mapping(map=mapping, bounds_postmap=[(-1, 1), (0, 1)])
square.add_hole(disk_L)
square.add_hole(disk_R)
# plot
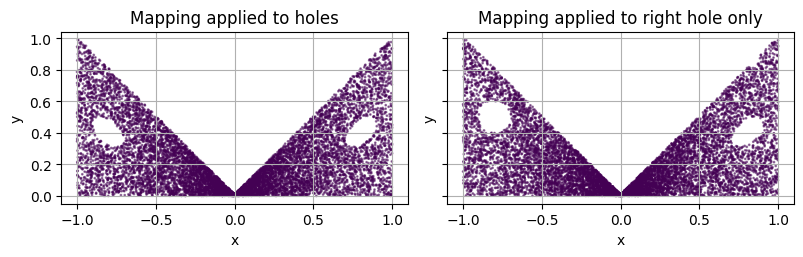
plot_2d(square, ax=axs[0])
axs[0].set_title('Mapping applied to holes')
# we can specify not to apply the mapping to a specific hole when adding it
square = simple_square()
disk_L, disk_R = simple_disks()
square.set_mapping(map=mapping, bounds_postmap=[(-1, 1), (0, 1)])
square.add_hole(disk_L, copy_mapping=False)
square.add_hole(disk_R, copy_mapping=True)
# plot
plot_2d(square, ax=axs[1])
axs[1].set_title('Mapping applied to right hole only')
# # Alternatively, we can specify not to apply the mapping to already existing holes,
# # but we will prefer to set the mapping first and then add the holes
# square = simple_square()
# disk_L, disk_R = simple_disks()
# square.add_hole(disk_L)
# square.set_mapping(map=mapping, bounds_postmap=[(-1, 1), (0, 1)], to_holes=False)
# square.add_hole(disk_R)
# plot2D(square, ax=axs[2])
# axs[2].set_title('Mapping applied to right hole only')
axs[2].remove()
fig.tight_layout()
plt.show()
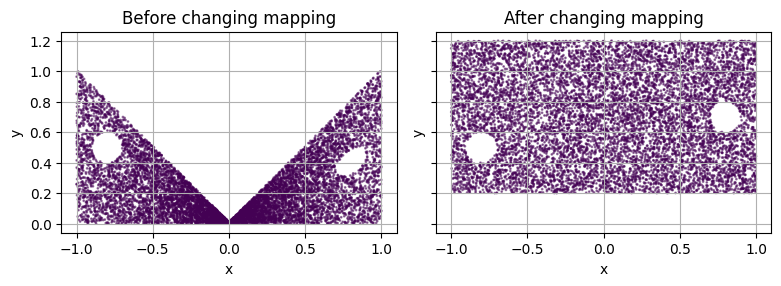
When changing the mapping of the domain (i.e. the domain was already mapped) with the flag to_holes set to False, the mapping will still be applied to previously mapped holes.
[9]:
fig, axs = plt.subplots(1, 2, figsize=(8, 4), sharex=True, sharey=True)
square = simple_square()
disk_L, disk_R = simple_disks()
# define a new mapping
mapping2 = Mapping.translate(torch.tensor([0, 0.2]))
# set the mapping for the square domain
square.set_mapping(map=mapping, bounds_postmap=[(-1, 1), (0, 1)])
# add the holes
square.add_hole(disk_L, copy_mapping=False) # left hole WITHOUT mapping
square.add_hole(disk_R) # right hole with mapping
# plot
plot_2d(square, ax=axs[0])
axs[0].set_title("Before changing mapping")
# apply the new mapping to the square domain
square.set_mapping(map=mapping2, bounds_postmap=[(-1, 1), (0.2, 1.2)], to_holes=False)
# plot
plot_2d(square, ax=axs[1])
axs[1].set_title("After changing mapping\n(previously mapped holes get new mapping)")
fig.tight_layout()
plt.show()
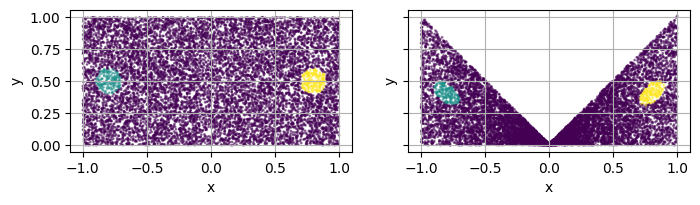
Create sub-domains¶
[10]:
fig, axs = plt.subplots(1, 2, figsize=(8, 4), sharex=True, sharey=True)
square = simple_square()
disk_L, disk_R = simple_disks()
square.add_subdomain(disk_L)
square.add_subdomain(disk_R)
plot_2d(square, ax=axs[0])
square.set_mapping(map=mapping, bounds_postmap=[(-1, 1), (0, 1)])
plot_2d(square, ax=axs[1])
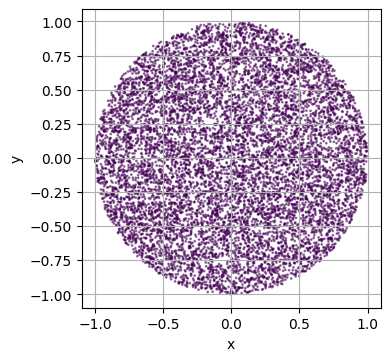
Defining your own Domains with a custom Signed Distance Function¶
A Signed distance function is simply a Callable with a threshold
[11]:
# import necessary libraries
from scimba_torch.domain.meshless_domain.base import VolumetricDomain
from scimba_torch.domain.sdf import SignedDistance
class CustomSDF(SignedDistance):
"""A custom SDF class that defines the signed distance function for unit 2D disk."""
def __init__(self):
super().__init__(
dim=2,
threshold=0.0, # threshold for inside points (sdf < -threshold)
threshold_out=0.0, # threshold for outside points, use for holes (sdf > threshold_out)
)
def __call__(self, pts: torch.Tensor) -> torch.Tensor:
"""Compute the signed distance function for the unit 2D disk."""
# pts: (N, dim) tensor
x, y = pts[:, 0], pts[:, 1]
return (torch.sqrt(x**2 + y**2) - 1.0)[:, None] # out: (N, 1) tensor
custom_domain = VolumetricDomain(
domain_type="CustomUnitDisk",
dim=2,
sdf=CustomSDF(),
bounds=[(-1, 1), (-1, 1)],
is_main_domain=True,
)
plot_2d(custom_domain)
/var/folders/zp/w4p46p_n0p52lgf7h0vpbrt00000gn/T/ipykernel_40294/3476767848.py:10: UserWarning: The domain does not have boundary domains
sampler = DomainSampler(domain)
Boundary Domains¶
TODO : implement default sampling with sdf when no bc domains are given ?
As you can see in the last example, our custom domain does not have any boundary domains…
In Scimba boundary domain (more generally called SurfacicDomains), are a “proper” domain (i.e. a VolumetricDomain) of dimension \(d\), mapped to dimension \(d+1\)
To declare such a domain, we hence need:
A
VolumetricDomain(calledparametric_domain)A
Mapping(calledsurface) that embeds ourparametric_domaininto higher dimension.
Let’s define the boundary of our unit disk.
[12]:
# import necessary libraries
from scimba_torch.domain.meshless_domain.base import SurfacicDomain
bc = SurfacicDomain(
domain_type="CustomUnitCircle",
parametric_domain=Segment1D((0, 2 * torch.pi)),
surface=Mapping(
from_dim=1,
to_dim=2,
map=lambda x: torch.stack([torch.cos(x[..., 0]), torch.sin(x[..., 0])], dim=-1),
),
)
# we then add the boundary domain to our custom domain
custom_domain.add_bc_domain(bc)
# we don't have a warning anymore :)
plot_2d(custom_domain)
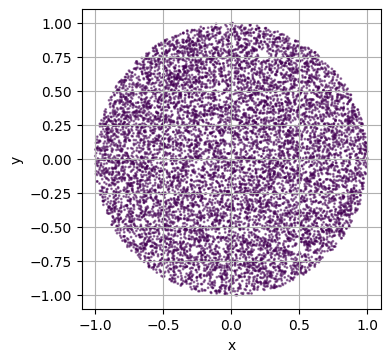
Let’s create a new plotting utility function to see what’s happening.
[13]:
# a utility function to plot a 2D domain and boundary points
def plot_2d_with_bc(domain, n=10_000, n_bc=1_000, ax=None):
"""Plot a 2D domain using a Domain sampler."""
cmap = plt.get_cmap("tab10_r")
# create the sampler
sampler = DomainSampler(domain)
# sample points
res = sampler.sample(n)
pts = res.x # coordinates of the sampled points
labels = res.labels # labels of the sampled points
# sample boundary points
res_bc, _ = sampler.bc_sample(n_bc)
pts_bc = res_bc.x # coordinates of the sampled boundary points
labels_bc = res_bc.labels # labels of the sampled boundary points
# convert to numpy for plotting
pts = pts.detach().cpu().numpy()
labels = labels.detach().cpu().numpy()
pts_bc = pts_bc.detach().cpu().numpy()
labels_bc = labels_bc.detach().cpu().numpy()
# generate the ax if not provided
if ax is None:
fig = plt.figure(figsize=(4, 4))
ax = fig.add_subplot(111, xlabel="x", ylabel="y", aspect="equal")
else:
fig = None
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.set_aspect("equal")
# offset the labels for boundary points
labels_bc += labels.max() + 1
max_label = max(labels.max(), labels_bc.max())
# plot the points
ax.scatter(
pts[:, 0],
pts[:, 1],
s=1,
c=labels,
alpha=0.5,
cmap=cmap,
vmin=0,
vmax=max_label,
)
# plot the boundary points
ax.scatter(
pts_bc[:, 0],
pts_bc[:, 1],
s=1,
c=labels_bc,
cmap=cmap,
vmin=0,
vmax=max_label,
alpha=0.5,
)
if fig is not None:
fig.tight_layout()
plt.show()
plot_2d_with_bc(custom_domain)
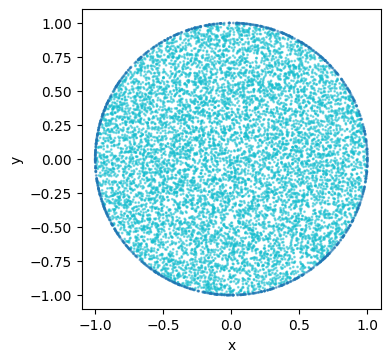
!Warning! Be aware that their is not check or whatsoever to see it the given boundary domain is really the boundary of the given domain.
Square2D domain implements full_bc_domain, which returns a list of SurfacicDomains that represent the boundary of a square.full_bc_domain.Complement on the mapping¶
Note that the Mapping can take extra arguments:
jacto declare the jacobian of the mapping (if not provided, it is computed using torchtorch.func.jacrev)invto be declared if the mapping is considered invertibleinv_jacfor the jacobian of the inverse (otherwise computed asjac)
sample_post_map.[14]:
from scimba_torch.integration.monte_carlo import SurfacicSampler, VolumetricSampler
# define a mapping function
def map(x):
"""A simple mapping function."""
return torch.stack([x[:, 0], x[:, 1]*(torch.abs(x[:, 0]) + 1e-2)], dim=1)
# define the inverse mapping function
def inv(y):
"""Inverse of the mapping function."""
return torch.stack([y[:, 0], y[:, 1]/(torch.abs(y[:, 0]) + 1e-2)], dim=1)
# define the mapping
mapping = Mapping(
from_dim=2,
to_dim=2,
map=map,
inv=inv
)
square = simple_square()
# set the mapping for the square domain
square.set_mapping(
map=mapping,
bounds_postmap=[(-1, 1), (0, 1)],
)
# create the sampler
sampler = VolumetricSampler(square)
# sample points
pts = sampler.sample_postmap(5_000)
pts = pts.detach().cpu().numpy()
# plot the points
fig, axs = plt.subplots(1, 2, figsize=(8, 4), sharex=True, sharey=True)
axs[0].scatter(pts[:, 0], pts[:, 1], s=1, c=torch.zeros(pts.shape[0]).numpy())
axs[0].set_aspect('equal')
plot_2d(square, ax=axs[1])
fig.tight_layout()
plt.show()
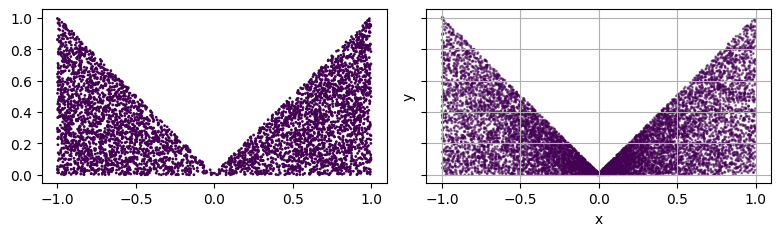
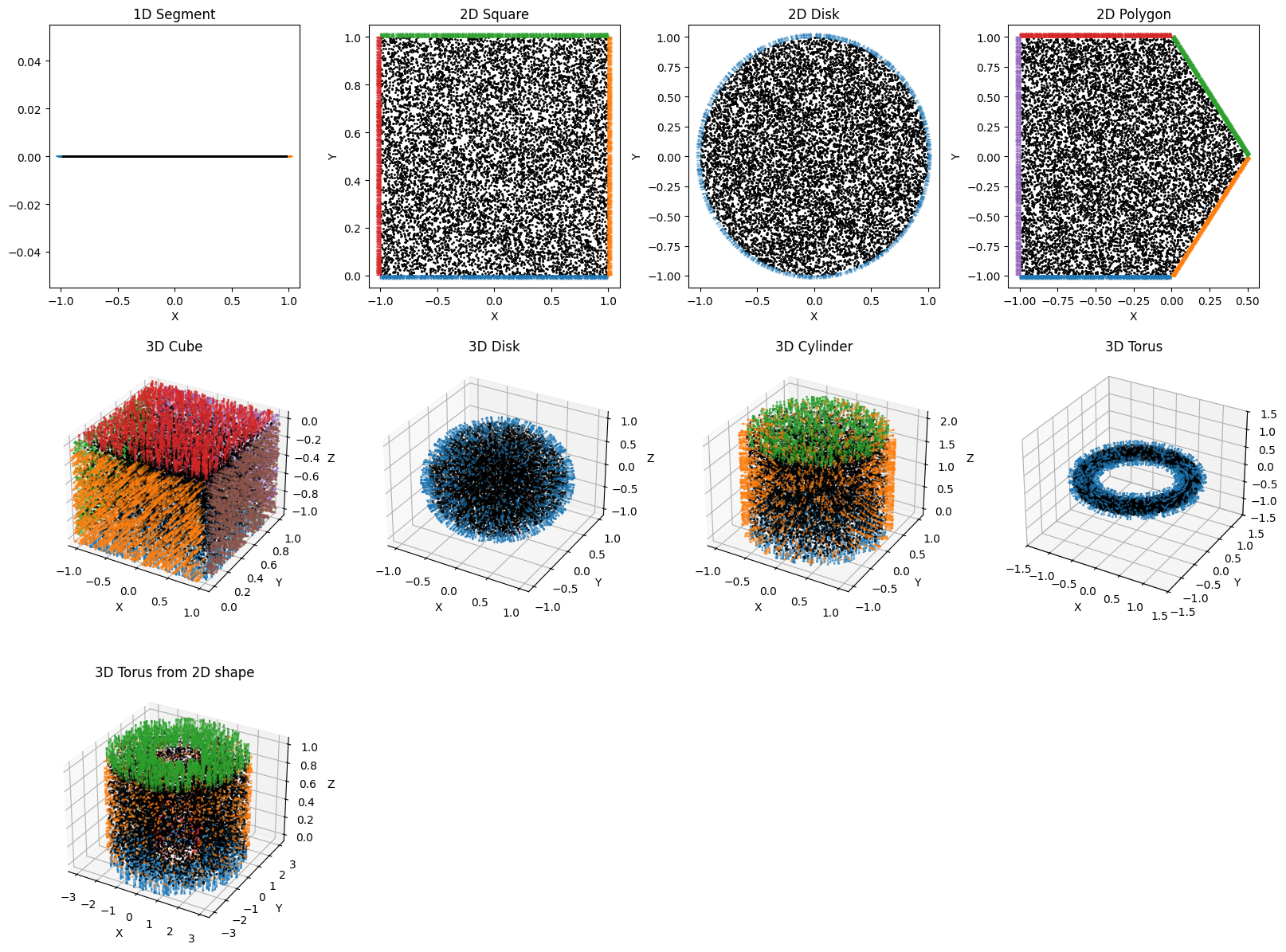
Available domain showcase¶
[15]:
# 1D domains
segment = Segment1D(
low_high=(-1, 1),
is_main_domain=True
)
# 2D domains
square = Square2D(
bounds=[(-1, 1), (0, 1)],
is_main_domain=True
)
disk = Disk2D(
center=(0, 0),
radius=1,
is_main_domain=True
)
polygon = Polygon2D(
# vertices of the polygon, they should be provided in counter-clockwise order
vertices=[[-1, -1], [0, -1], [.5, 0], [0, 1], [-1, 1]],
is_main_domain=True
)
# 3D domains
cube = Cube3D(
bounds=[(-1, 1), (0, 1), (-1, 0)],
is_main_domain=True
)
disk3d = Disk3D(
center=(0, 0, 0),
radius=1,
is_main_domain=True
)
cylinder = Cylinder3D(
radius=1,
length=2,
is_main_domain=True
)
torus = Torus3D(
center=(0, 0, 0),
radius=1,
tube_radius=0.2,
is_main_domain=True
)
torus_from2d = TorusFrom2DVolume(
base_volume=square,
radius=2,
is_main_domain=True
)
[16]:
# import necessary libraries
from scimba_torch.integration.monte_carlo import VolumetricSampler
# utility function to sample and plot points in a domain
def plot(dict_dom2ax, n=10_000, n_bc=1000):
for domain, ax in dict_dom2ax.items():
# create a sampler for the domain
sampler = VolumetricSampler(domain)
bc_sampler = [SurfacicSampler(bc) for bc in domain.full_bc_domain()]
# sample N points
pts = sampler.sample(n)
bc_pts = [
bc_sampler[i].sample(n_bc, compute_normals=True)
for i in range(len(bc_sampler))
]
# convert to numpy for plotting (& normalize normals)
pts = pts.detach().numpy()
bc_pts = [
(bc, 0.1 * n / torch.linalg.norm(n, axis=-1, keepdims=True))
for bc, n in bc_pts
]
bc_pts = [(bc[0].detach().numpy(), bc[1].detach().numpy()) for bc in bc_pts]
def get_pts(pts):
if domain.dim == 1:
return pts[:, 0], [0] * len(pts)
elif domain.dim == 2:
return pts[:, 0], pts[:, 1]
elif domain.dim == 3:
return pts[:, 0], pts[:, 1], pts[:, 2]
else:
raise ValueError("Domain dimension not supported for plotting.")
ax.scatter(*get_pts(pts), color="k", s=1) # plot the sampled points
for i, (bc, nn) in enumerate(bc_pts):
ax.scatter(
*get_pts(bc), s=1, c=f"C{i}", alpha=0.2
) # plot the boundary points
ax.quiver(
*get_pts(bc), *get_pts(nn), color=f"C{i}", alpha=0.5
) # plot the boundary normals
fig = plt.figure(figsize=(16, 12))
dict_dom2ax = {
segment: fig.add_subplot(3, 4, 1, xlabel="X", title="1D Segment"),
square: fig.add_subplot(3, 4, 2, xlabel="X", ylabel="Y", title="2D Square"),
disk: fig.add_subplot(3, 4, 3, xlabel="X", ylabel="Y", title="2D Disk"),
polygon: fig.add_subplot(3, 4, 4, xlabel="X", ylabel="Y", title="2D Polygon"),
cube: fig.add_subplot(
3, 4, 5, projection="3d", xlabel="X", ylabel="Y", zlabel="Z", title="3D Cube"
),
disk3d: fig.add_subplot(
3, 4, 6, projection="3d", xlabel="X", ylabel="Y", zlabel="Z", title="3D Disk"
),
cylinder: fig.add_subplot(
3,
4,
7,
projection="3d",
xlabel="X",
ylabel="Y",
zlabel="Z",
title="3D Cylinder",
),
torus: fig.add_subplot(
3,
4,
8,
projection="3d",
xlabel="X",
ylabel="Y",
zlabel="Z",
xlim=(-1.5, 1.5),
ylim=(-1.5, 1.5),
zlim=(-1.5, 1.5),
title="3D Torus",
),
torus_from2d: fig.add_subplot(
3,
4,
9,
projection="3d",
xlabel="X",
ylabel="Y",
zlabel="Z",
title="3D Torus from 2D shape",
),
}
# plot the domains
plot(dict_dom2ax)
fig.tight_layout()
plt.show()
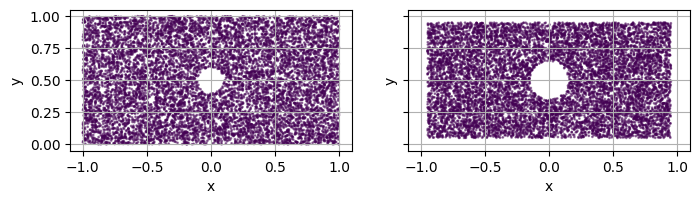
Using the threshold to avoid sampling near the boundary¶
set_threshold on a main domain.threshold (resp. threshold_out) of the main domain (resp. holes).domain.sdf.threshold for the domain and hole.sdf.threshold_out for each hole.TODO : “uniformize” sdf of domains ?
[17]:
fig, axs = plt.subplots(1, 2, figsize=(8, 4), sharex=True, sharey=True)
square = simple_square()
hole = Disk2D(center=(0, 0.5), radius=0.1, is_main_domain=False)
square.add_hole(hole)
# plot with default threshold
plot_2d(square, ax=axs[0])
# set a threshold to avoid sampling near the boundary
square.set_threshold(0.05)
plot_2d(square, ax=axs[1])