Scimba basics II: Natural gradient preconditioners¶
In the previous tutorials, we used a PINN fit through a natural gradient descent algorithm to the solution of a 2D laplacian problem.
Scimba implements several natural gradient preconditioning approaches for gradient descent algorithms: Energy Natural Gradient (ENG), ANaGRAM, Sketchy Natural Gradient (SNG) and Nyström Natural Gradient (NNG).
ENG and Anagram can be used for the task of projecting a function onto an approximation space while all can be used for PINNs.
This tutorial demonstrates further the use of preconditioners for PINNs on the 2D laplacian example, and for projectors. A following tutorial explains how to model a pde so that it can be used with a natural gradient preconditioned PINN.
Let us first define the problem we consider, and solve it with a not preconditioned PINN for the reference.
Approximation with a not preconditioned PINN¶
[1]:
import torch
from scimba_torch.approximation_space.nn_space import NNxSpace
from scimba_torch.domain.meshless_domain.domain_2d import Square2D
from scimba_torch.integration.monte_carlo import DomainSampler, TensorizedSampler
from scimba_torch.integration.monte_carlo_parameters import UniformParametricSampler
from scimba_torch.neural_nets.coordinates_based_nets.mlp import GenericMLP
from scimba_torch.physical_models.elliptic_pde.laplacians import (
Laplacian2DDirichletStrongForm,
)
from scimba_torch.utils.scimba_tensors import LabelTensor
domain_x = Square2D([(0.0, 1), (0.0, 1)], is_main_domain=True)
domain_mu = [(0.5, 1.0)]
sampler = TensorizedSampler(
[DomainSampler(domain_x), UniformParametricSampler(domain_mu)]
)
def f_rhs(xs: LabelTensor, ms: LabelTensor) -> torch.Tensor:
x, y = xs.get_components()
mu = ms.get_components()
pi = torch.pi
return 2 * (2.0 * mu * pi) ** 2 * torch.sin(2.0 * pi * x) * torch.sin(2.0 * pi * y)
def f_bc(xs: LabelTensor, ms: LabelTensor) -> torch.Tensor:
x, _ = xs.get_components()
return x * 0.0
torch.manual_seed(0)
space = NNxSpace(
1, # the output dimension
1, # the parameter's space dimension
GenericMLP, # the type of neural network
domain_x, # the geometric domain
sampler, # the sampler
layer_sizes=[16, 16], # the size of intermediate layers
)
pde = Laplacian2DDirichletStrongForm(space, f=f_rhs, g=f_bc)
We define a PinnsElliptic object to approximate the solution of pde.
Notice the use of keyword argument bc_weight to scale the loss induced by the boundary condition relatively to the loss of the residual inside the domain.
[2]:
from scimba_torch.numerical_solvers.elliptic_pde.pinns import PinnsElliptic
pinn = PinnsElliptic(pde, # the pde to be solved
bc_type="weak", # use explicitly the boundary conditions
bc_weight=50.) # the weight of the loss of the boundary condition
The next step is to solve the PINN, that is to find the parameters of the approximation space (here a multi-layer perceptron) to minimize the residual inside the domain and the boundary condition, here Dirichlet.
At each step, this requires to sample the interior and the boundaries of the considered domain \(\Omega\). Below, we time 1000 steps of the Adam optimizer.
[3]:
import timeit
pinn.solve(
epochs=1000, #number of steps of optimization
n_collocation=1000, #number of sample points inside the domain
n_bc_collocation=1000, #number of sample points on the boundary
)
print("ENG training time: %.2f s"%pinn.training_time)
print("best loss: %.2e"%pinn.best_loss)
Training: 100%|||||||||||||||| 1000/1000[00:04<00:00] , loss: 6.0e+02 -> 1.7e+01
ENG training time: 5.06 s
best loss: 1.73e+01
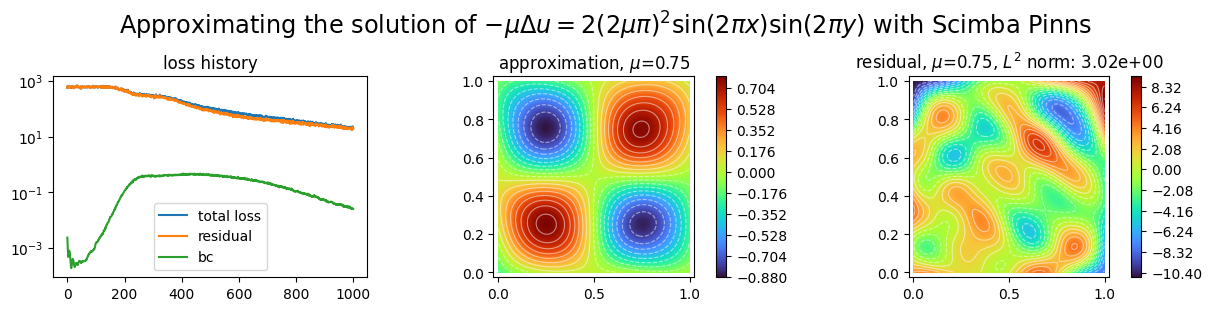
Let us plot the obtained approximation, the residual inside the domain and the history of the loss variation during the optimization.
[4]:
import matplotlib.pyplot as plt
from scimba_torch.plots.plots_nd import plot_abstract_approx_spaces
plot_abstract_approx_spaces(
pinn.space, # the approximation space
domain_x, # the spatial domain
domain_mu, # the parameter's domain
loss=pinn.losses, # optional plot of the loss: the losses
residual=pinn.pde, # optional plot of the residual of the pde
draw_contours=True, # plotting isolevel lines
n_drawn_contours=20, # number of isolevel lines,
title=(
"Approximating the solution of "
r"$- \mu\Delta u = 2(2\mu\pi)^2 \sin(2\pi x) \sin(2\pi y)$"
" with Scimba Pinns"
),
)
plt.show()
Approximation with preconditioned PINNs¶
Scimba implements four natural gradient preconditioners for PINNs.
They all rely on computing an approximate solution of \(Gx=y\) where \(G\) (a square matrix of size \(p\times p\) where \(p\) is the number of degrees of freedom of the NN) is the so called energy Gram matrix and \(y\) (a vector of size \(p\)) is the current gradient of the loss with respect to the degrees of freedom of the NN; they differ in how the approximate solution is computed.
PINNs with natural gradient preconditioning are implemented by the NaturalGradientPinnsElliptic class for stationary problems, and by the NaturalGradientTemporalPinns class for time dependent problems.
By default, these two classes implement ENG preconditioning; other approaches are obtained with the keyword argument ng_algo that can take the values:
“ANaGRAM”,
“SNG” or “sketchy”, and
“NNG” or Nyström.
Energy Natural Gradient preconditioning¶
[5]:
torch.manual_seed(0)
ENGspace = NNxSpace(1, 1, GenericMLP, domain_x, sampler, layer_sizes=[16, 16])
ENGpde = Laplacian2DDirichletStrongForm(ENGspace, f=f_rhs, g=f_bc)
By default, NaturalGradientPinnsElliptic use ENG preconditioning.
Notice the use of the matrix_regularization argument when instantiating the pinn. It is set to \(10^{-6}\) by default. In your examples, play around with this parameter as preconditioning with ENG is sensitive to this parameter.
[6]:
from scimba_torch.numerical_solvers.elliptic_pde.pinns import (
NaturalGradientPinnsElliptic,
)
ENGpinn = NaturalGradientPinnsElliptic(
ENGpde,
bc_type="weak",
bc_weight=50.,
matrix_regularization=1e-6
)
Use SGD algorithm together with a line search for adaptive choice of learning rate.
[7]:
ENGpinn.solve(epochs=100,
n_collocation=1000,
n_bc_collocation=1000)
print("ENG training time: %.2f s"%ENGpinn.training_time)
print("best loss: %.2e"%ENGpinn.best_loss)
Training: 100%|||||||||||||||||| 100/100[00:02<00:00] , loss: 6.0e+02 -> 8.9e-05
ENG training time: 2.37 s
best loss: 8.87e-05
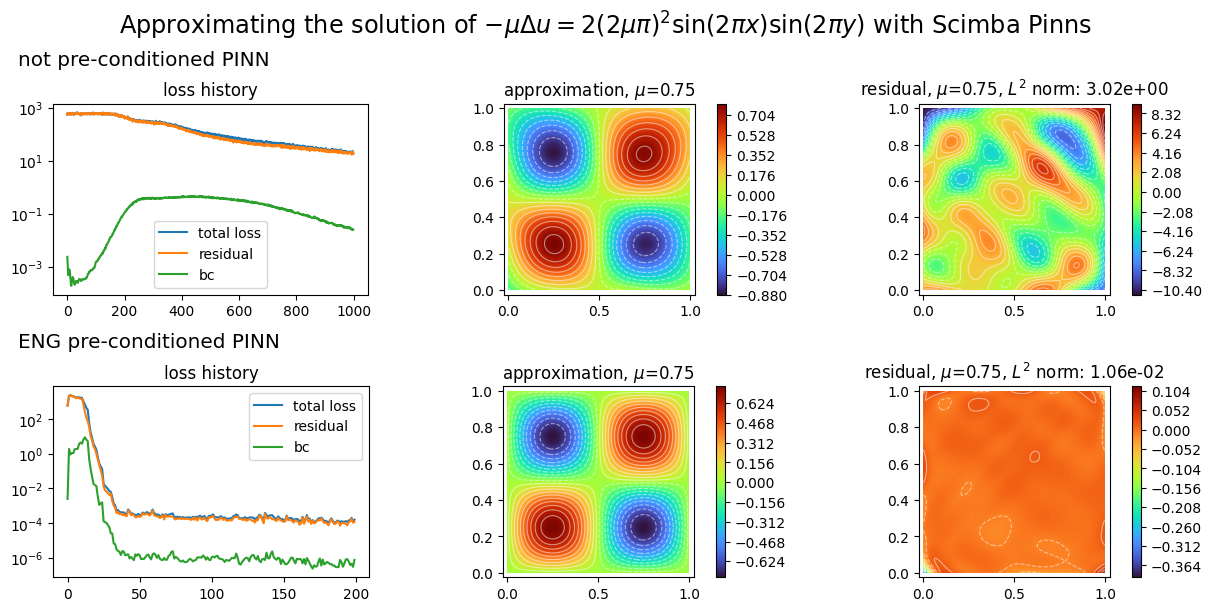
We plot the two PINNs with:
[8]:
plot_abstract_approx_spaces(
(pinn.space, ENGpinn.space), # the approximation spaces
domain_x, # the spatial domain
domain_mu, # the parameter's domain
loss=(pinn.losses, ENGpinn.losses), # optional plot of the loss: the losses
residual=(pinn.pde, ENGpinn.pde), # optional plot of the residual of the pde
draw_contours=True, # plotting isolevel lines
n_drawn_contours=20, # number of isolevel lines,
title=(
"Approximating the solution of "
r"$- \mu\Delta u = 2(2\mu\pi)^2 \sin(2\pi x) \sin(2\pi y)$"
" with Scimba Pinns"
),
titles=("not preconditioned PINN", "ENG preconditioned PINN"),
)
plt.show()
ANaGRAM preconditioning¶
In order to demonstrate ANaGRAM use, we define new approximation space and PDE:
[9]:
torch.manual_seed(0)
ANAspace = NNxSpace(1, 1, GenericMLP, domain_x, sampler, layer_sizes=[16, 16])
ANApde = Laplacian2DDirichletStrongForm(ANAspace, f=f_rhs, g=f_bc)
On uses the keyword argument ng_algo to choose the gradient descent preconditioning algorithm.
Notice the use below of the svd_threshold argument when creating the PINN object. The success of the optimization is very sensitive to this parameter; its default value of \(10^{-6}\) might be a very bad choice for some examples.
[10]:
ANApinn = NaturalGradientPinnsElliptic(
ANApde, bc_type="weak", bc_weight=50.0, ng_algo="ANaGRAM", svd_threshold=0.1
)
ANApinn.solve(epochs=100, n_collocation=1000, n_bc_collocation=1000, bool_time=True)
print("ANaGRAM training time: %.2f s"%ANApinn.training_time)
print("best loss: %.2e"%ANApinn.best_loss)
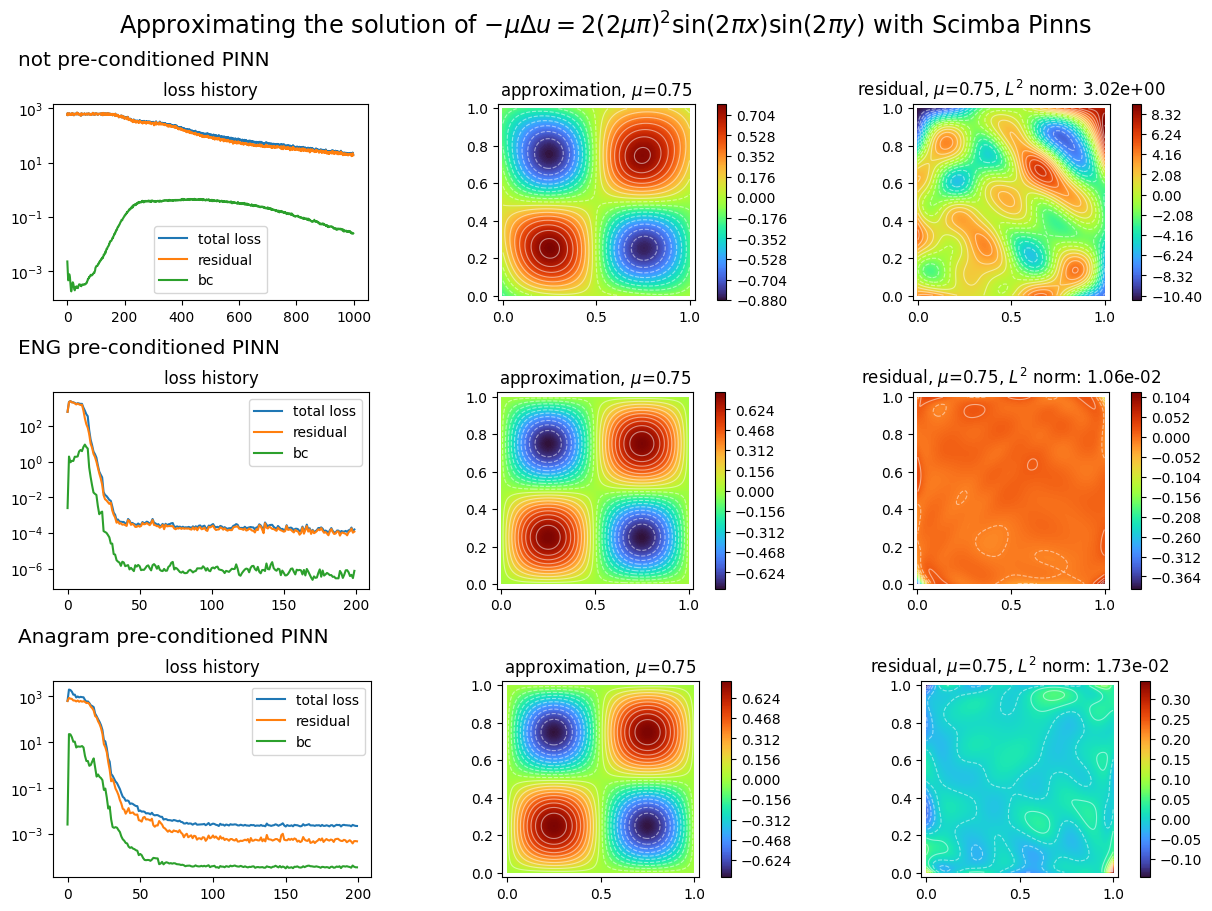
plot_abstract_approx_spaces(
(pinn.space, ENGpinn.space, ANApinn.space), # the approximation space
domain_x, # the spatial domain
domain_mu, # the parameter's domain
loss=(
pinn.losses,
ENGpinn.losses,
ANApinn.losses,
), # optional plot of the loss: the losses
residual=(
pinn.pde,
ENGpinn.pde,
ANApinn.pde,
), # optional plot of the residual of the pde
draw_contours=True, # plotting isolevel lines
n_drawn_contours=20, # number of isolevel lines,
title=(
"Approximating the solution of "
r"$- \mu\Delta u = 2(2\mu\pi)^2 \sin(2\pi x) \sin(2\pi y)$"
" with Scimba Pinns"
),
titles=(
"not preconditioned PINN",
"ENG preconditioned PINN",
"ANaGRAM preconditioned PINN",
),
)
plt.show()
Training: 100%|||||||||||||||||| 100/100[00:05<00:00] , loss: 6.0e+02 -> 2.4e-03
ANaGRAM training time: 5.39 s
best loss: 2.39e-03
Sketchy Natural Gradient preconditioning¶
We illustrate below the use of Sketchy Natural Gradient preconditioning.
[11]:
torch.manual_seed(0)
SNGspace = NNxSpace(1, 1, GenericMLP, domain_x, sampler, layer_sizes=[16, 16])
SNGpde = Laplacian2DDirichletStrongForm(SNGspace, f=f_rhs, g=f_bc)
SNGpinn = NaturalGradientPinnsElliptic(
SNGpde, bc_type="weak", bc_weight=50.0, ng_algo="SNG", tol=1e-7
)
SNGpinn.solve(epochs=100, n_collocation=1000, n_bc_collocation=1000)
print("SNG training time: %.2f s"%SNGpinn.training_time)
print("best loss: %.2e"%SNGpinn.best_loss)
Training: 100%|||||||||||||||||| 100/100[00:02<00:00] , loss: 6.0e+02 -> 9.2e-03
SNG training time: 2.54 s
best loss: 9.20e-03
Nyström Natural Gradient preconditioning¶
We illustrate below the use of Nyström Natural Gradient preconditioning.
[12]:
torch.manual_seed(0)
NNGspace = NNxSpace(1, 1, GenericMLP, domain_x, sampler, layer_sizes=[16, 16])
NNGpde = Laplacian2DDirichletStrongForm(NNGspace, f=f_rhs, g=f_bc)
NNGpinn = NaturalGradientPinnsElliptic(
NNGpde, bc_type="weak", bc_weight=50.0, ng_algo="NNG", eps=1e-12
)
NNGpinn.solve(epochs=100, n_collocation=1000, n_bc_collocation=1000)
print("NNG training time: %.2f s"%NNGpinn.training_time)
print("best loss: %.2e"%NNGpinn.best_loss)
Training: 100%|||||||||||||||||| 100/100[00:03<00:00] , loss: 6.0e+02 -> 4.5e-04
NNG training time: 3.13 s
best loss: 4.47e-04
Nyström Natural Gradient can be done in a matrix free way:
[13]:
torch.manual_seed(0)
NNGMFspace = NNxSpace(1, 1, GenericMLP, domain_x, sampler, layer_sizes=[16, 16])
NNGMFpde = Laplacian2DDirichletStrongForm(NNGMFspace, f=f_rhs, g=f_bc)
NNGMFpinn = NaturalGradientPinnsElliptic(
NNGMFpde, bc_type="weak", bc_weight=50.0, ng_algo="NNG", eps=1e-12, matrix_free=True
)
NNGMFpinn.solve(epochs=100, n_collocation=1000, n_bc_collocation=1000)
print("NNGMF training time: %.2f s"%NNGMFpinn.training_time)
print("best loss: %.2e"%NNGMFpinn.best_loss)
Training: 100%|||||||||||||||||| 100/100[00:25<00:00] , loss: 6.0e+02 -> 5.6e-04
NNGMF training time: 25.72 s
best loss: 5.61e-04
We explain in this tutorial how to write a model for which natural gradient preconditioners can be used.
preconditioned projectors¶
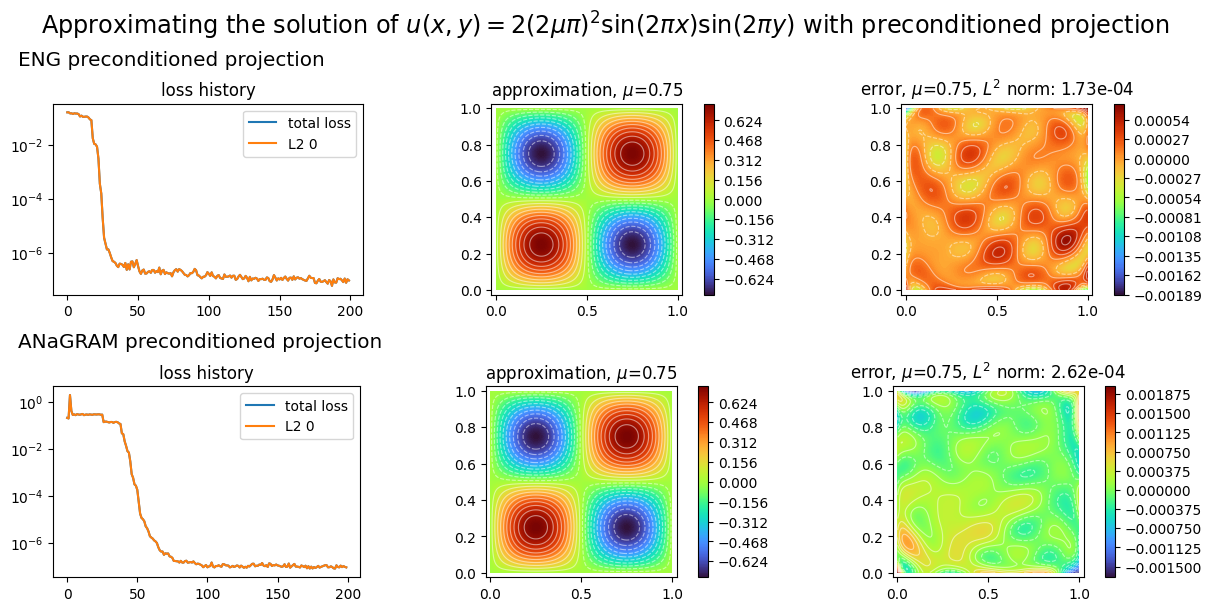
To finish with, we show how to project a known function on an approximation space with preconditioned gradient descent optimization. We project the exact solution the 2D laplacian problem considered so far.
[14]:
from scimba_torch.numerical_solvers.collocation_projector import (
NaturalGradientProjector,
)
def exact_sol(xs: LabelTensor, ms: LabelTensor) -> torch.Tensor:
x, y = xs.get_components()
mu = ms.get_components()
pi = torch.pi
return mu * torch.sin(2.0 * pi * x) * torch.sin(2.0 * pi * y)
ENGspace = NNxSpace(1, 1, GenericMLP, domain_x, sampler, layer_sizes=[16, 16])
ENGprojector = NaturalGradientProjector(ENGspace, exact_sol)
ENGprojector.solve(epochs=200, n_collocation=2000)
print("ENG training time: %.2f s"%ENGprojector.training_time)
print("best L2 error: %.2e"%ENGprojector.best_loss)
ANAspace = NNxSpace(1, 1, GenericMLP, domain_x, sampler, layer_sizes=[16, 16])
ANAprojector = NaturalGradientProjector(ANAspace, exact_sol, ng_algo="ANaGRAM", svd_threshold=5e-3)
ANAprojector.solve(epochs=200, n_collocation=2000)
print("ANaGRAM training time: %.2f s"%ANAprojector.training_time)
print("best L2 error: %.2e"%ANAprojector.best_loss)
Training: 100%|||||||||||||||||| 200/200[00:01<00:00] , loss: 1.6e-01 -> 5.8e-08
ENG training time: 1.70 s
best L2 error: 5.84e-08
Training: 100%|||||||||||||||||| 200/200[00:07<00:00] , loss: 2.0e-01 -> 8.1e-08
ANaGRAM training time: 7.94 s
best L2 error: 8.07e-08
[15]:
plot_abstract_approx_spaces(
(ENGprojector.space, ANAprojector.space), # the approximation space
domain_x, # the spatial domain
domain_mu, # the parameter's domain
# optional plot of the loss: the losses
loss=(ENGprojector.losses, ANAprojector.losses),
error=exact_sol, # plot the error w.r.t. the exact solution
draw_contours=True, # plotting isolevel lines
n_drawn_contours=20, # number of isolevel lines,
title=(
"Approximating the solution of "
r"$u(x,y) = 2(2\mu\pi)^2 \sin(2\pi x) \sin(2\pi y)$"
" with preconditioned projection"
),
titles=("ENG preconditioned projection", "ANaGRAM preconditioned projection"),
)
plt.show()
[ ]: